BDB-Lab members involved
Persistence of High-Risk Antimicrobial Resistance Genes in Extracellular DNA Along an Urban Wastewater-River Continuum
Persistence of High-Risk Antimicrobial Resistance Genes in Extracellular DNA Along an Urban Wastewater-River Continuum
by John Paul Makumbi, Samuel K. Leareng, Oliver K. Bezuidt, Luis Pedro Coelho, Thulani P. Makhalanyane
Abstract
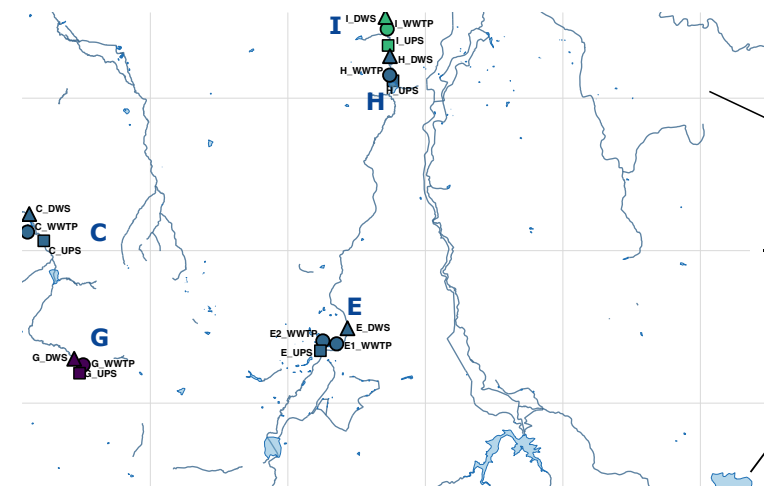
Inadequate wastewater treatment can drive the spread of antimicrobial resistance (AMR), posing risks to ecosystems and human health. Extracellular DNA (exDNA) plays a key role in AMR persistence by stabilizing antimicrobial resistance genes (ARGs) in the environment and facilitating horizontal gene transfer (HGT). However, its taxonomic structure and precise influence on AMR ecology remain poorly understood, especially in African aquatic systems. We investigated exDNA-associated resistomes across nine South African wastewater treatment plants (WWTPs) and receiving rivers, comparing single-stage activated sludge process (ASP29 only) and combined ASP-biofilter systems. We found that exDNA harbours high-risk ARGs, including potentially mobile genes conferring resistance to last-resort antibiotics. ARGs were enriched in effluents and downstream rivers, especially where exDNA removal was inefficient. Surprisingly, ARGs were also abundant upstream of WWTPs, suggesting riverine reservoirs of exDNA-associated AMR. ARGs were mainly associated with Pseudomonadota and Bacteroidota phyla, suggesting that exDNA is an ecologically distinct AMR reservoir dominated by a few key taxa. These findings highlight the need to integrate exDNA into AMR surveillance and highlight its broader ecological role in shaping microbial evolution and adaptation in freshwater environments.
Full text: https://doi.org/10.1016/j.celrep.2026.117128
Copyright (c) 2018–2026. Luis Pedro Coelho and other group members. All rights reserved.

